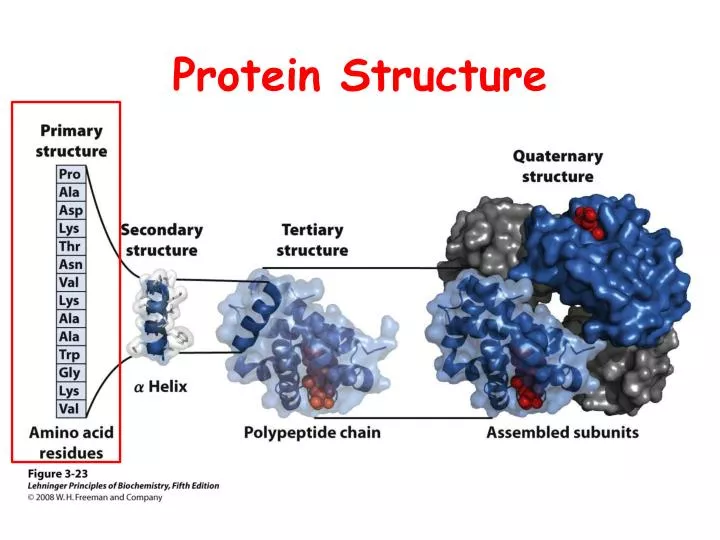
I think such a resource would be more of a hindrance than an asset to the scientific community. If there was a database of predicted secondary structures, people would likely take them for granted (make the equation prediction = fact) which would be quite "unscientific". Alternatively, click on the launch icon to open the advanced (full feature) version of iCn3D, NCBI's web-based 3D structure viewer, in a separate window. Scroll to the molecular graphic section and click on the spin icon to load an interactive view of the structure within the web page. Also they change each time the algorithms are improved. Enter the PDB code in the search box and press the Go button. The results may differ from one prediction method to another. 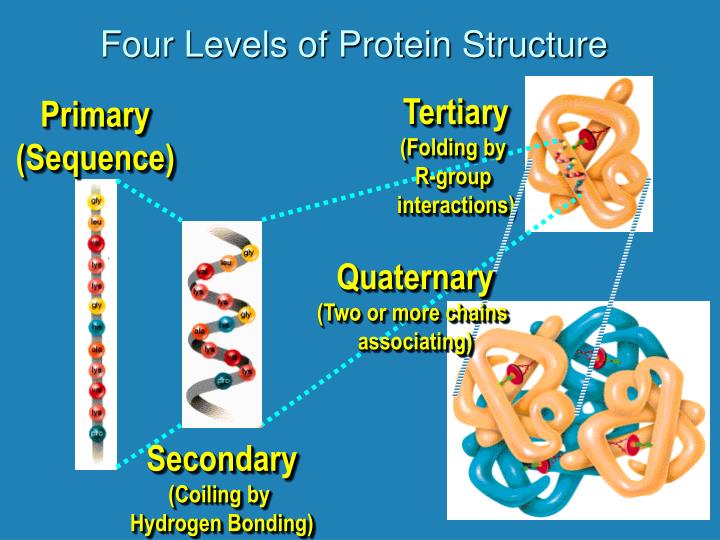
They're "educated guesses" that are often correct, but sometimes wrong. And I use them quite frequently: they are extremely useful! But as far as I know, no prediction is accepted as fact. In those cases, it is not only the secondary structures but also the tertiary structures (with the caveat that the crystal structure of a protein does not prove "all" states that a protein may take in real "dynamic" physiological conditions).įor all those proteins that have not been crystalized, then we can only rely on predictions. That may be the case for secondary structures of proteins, but only in the case where the said proteins have been crystalized. I'm not sure that makes much sense, let me explain my point of view (which may not be that of everyone): most often, the data that is found in databases is the "state of knowledge" of the described object, based on experimentation. You suggest that there should be a Secondary Structure Database. I think you found the best answer yourself: use a predictor! There are several out there.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
